Deep learning in genomics is revolutionizing how we analyze and interpret vast amounts of biological data, providing opportunities for groundbreaking discoveries. At LEARNS.EDU.VN, we aim to provide an accessible and comprehensive overview of how deep learning algorithms are transforming genomic research, offering solutions to complex analytical challenges and unlocking new insights. Discover advanced pattern recognition and predictive modeling techniques.
1. Introduction to Deep Learning in Genomics
Genomics, the study of genomes, has been transformed by the advent of high-throughput sequencing technologies. These technologies generate massive datasets that are complex and require advanced computational techniques for analysis. Traditional bioinformatics approaches often fall short in capturing the intricate patterns and relationships within these datasets. This is where deep learning, a subset of machine learning, comes into play.
Deep learning models, inspired by the structure and function of the human brain, are capable of learning complex representations from data. These models consist of multiple layers of interconnected nodes (neurons) that process information in a hierarchical manner. Each layer learns to extract increasingly abstract features from the input data, allowing the model to capture intricate patterns and relationships.
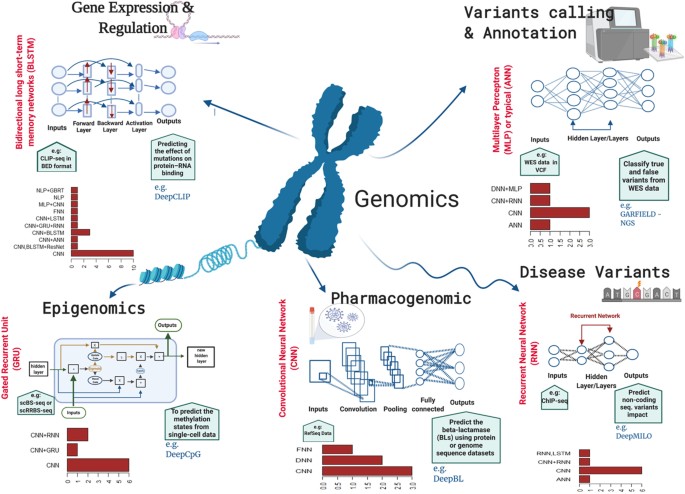
In the context of genomics, deep learning has been applied to a wide range of tasks, including:
- Variant calling and annotation
- Disease variant prediction
- Gene expression and regulation analysis
- Epigenomics
- Pharmacogenomics
1.1. The Role of High-Throughput Data
The ability to generate high-throughput data is central to the application of deep learning in genomics. High-throughput technologies, such as next-generation sequencing (NGS), allow researchers to rapidly and cost-effectively sequence DNA and RNA. This has led to an explosion of genomic data, providing the raw material needed to train deep learning models.
High-throughput data is characterized by its:
- Volume: Massive amounts of data are generated, often terabytes or even petabytes in size.
- Velocity: Data is generated at a rapid pace, requiring efficient processing and analysis techniques.
- Variety: Data comes in various forms, including DNA sequences, RNA sequences, gene expression levels, and epigenetic marks.
- Veracity: Data may contain errors and biases, requiring careful quality control and preprocessing.
1.2. Deep Learning Algorithms for Sophisticated Predictions
Deep learning algorithms excel at making sophisticated predictions from complex data. These algorithms can learn to identify subtle patterns and relationships that are not apparent to traditional methods. This capability is particularly valuable in genomics, where the underlying biological processes are often complex and poorly understood.
Common deep learning algorithms used in genomics include:
- Convolutional Neural Networks (CNNs): CNNs are particularly well-suited for analyzing sequence data, such as DNA and RNA sequences. They can learn to identify motifs and patterns within the sequences that are associated with specific biological functions.
- Recurrent Neural Networks (RNNs): RNNs are designed to handle sequential data, such as time series data or text. In genomics, RNNs can be used to analyze gene expression data over time or to model the interactions between genes.
- Deep Neural Networks (DNNs): DNNs are general-purpose deep learning models that can be applied to a wide range of tasks. In genomics, DNNs can be used to integrate data from multiple sources and to make predictions about complex biological processes.
- Generative Adversarial Networks (GANs): GANs are used for data augmentation and generation, helping to improve the robustness and accuracy of other deep learning models.
- Transformers: Transformers, with their attention mechanisms, can handle long-range dependencies in genomic sequences, which is crucial for understanding regulatory elements.
1.3. Transforming Big Biological Data
The ultimate goal of applying deep learning in genomics is to transform big biological data into new knowledge and novel findings. By analyzing large genomic datasets, deep learning models can help researchers:
- Identify disease-causing genes
- Predict drug response
- Understand gene regulation
- Develop new diagnostic and therapeutic strategies
LEARNS.EDU.VN is committed to providing resources and education to help researchers and students leverage the power of deep learning in genomics. Understanding these concepts will help unlock solutions for personalized medicine, drug discovery, and understanding complex biological systems.
2. Variant Calling and Annotation using Deep Learning
Variant calling and annotation are essential steps in genomic analysis. Variant calling involves identifying differences between an individual’s DNA sequence and a reference genome. Annotation involves providing information about the potential functional consequences of these variants. Deep learning is enhancing the accuracy and efficiency of these processes.
2.1. The Importance of NGS in Personalized Medicine
Next-generation sequencing (NGS) technologies, including whole-genome sequencing (WGS) and whole-exome sequencing (WES), are at the forefront of personalized medicine. These technologies enable the rapid and cost-effective sequencing of an individual’s genome, providing a wealth of information about their genetic makeup.
NGS plays a crucial role in:
- Identifying genetic variations associated with disease
- Predicting an individual’s response to drugs
- Guiding treatment decisions
- Monitoring disease progression
The ability to sequence thousands of human genomes has become routine, making personalized medicine a reality.
2.2. Challenges in High-Throughput Sequencing
Despite the advances in NGS technologies, several challenges remain. These include:
- Error rates: NGS procedures are prone to technical and bioinformatics errors, which can lead to false-positive variant calls.
- Computational problems: The massive amounts of data generated by NGS require sophisticated bioinformatics tools for processing and analysis.
- Variant discovery: Some actual variants may be missed due to limitations in sequencing coverage and bioinformatics algorithms.
These challenges highlight the need for improved variant calling and annotation methods.
2.3. Overcoming Limitations with Deep Learning Tools
Deep learning tools have emerged as a promising solution to overcome the limitations of traditional variant calling pipelines. By learning complex patterns in the data, deep learning models can improve the accuracy and sensitivity of variant calling.
Some notable deep learning tools for variant calling and annotation include:
- DeepVariant: Developed by Google, DeepVariant treats variant calling as an image classification task. It converts the mapped sequencing datasets into images and uses a convolutional neural network (CNN) to identify genetic variants.
- DeepSV: DeepSV is designed to predict long genomic deletions from sequencing read images. It processes BAM or VCF files as input and outputs the results in VCF format.
- GARFIELD-NGS: GARFIELD-NGS is a genomic variant filtering tool that relies on a multilayer perceptron (MLP) algorithm to identify true and false variants in exome sequencing datasets.
- Clairvoyante: Clairvoyante is a variant caller designed to utilize long-read sequencing data generated from single-molecule sequencing (SMS) technologies. It predicts variant type, zygosity, allele alternative, and Indel length.
- Intelli-NGS: Intelli-NGS is an artificial neural network (ANN)-based variant caller that identifies true and false variants from Ion Torrent sequencer data.
2.4. Case Study: DeepVariant
DeepVariant, developed by Google, exemplifies the application of deep learning in variant calling. DeepVariant uses a convolutional neural network (CNN) to classify genetic variants from NGS data. It treats the mapped sequencing datasets as images and converts the variant calls into image classification tasks.
Key features of DeepVariant include:
- High accuracy: DeepVariant has been shown to improve the accuracy of single-nucleotide variant (SNV) and Indel detections compared to traditional variant callers.
- Generalizability: DeepVariant can be applied to a wide range of sequencing datasets and variant types.
- Open-source: DeepVariant is available as an open-source tool, making it accessible to researchers and clinicians.
2.5. Implementing Deep Learning in Variant Calling
Implementing deep learning in variant calling involves several steps:
- Data preparation: Preprocess the NGS data to remove errors and biases.
- Model selection: Choose an appropriate deep learning model for the task, such as DeepVariant or Clairvoyante.
- Training: Train the deep learning model on a large dataset of known variants.
- Validation: Validate the performance of the deep learning model on an independent dataset.
- Deployment: Integrate the deep learning model into a variant calling pipeline.
LEARNS.EDU.VN provides resources and courses to help researchers and students implement deep learning in variant calling and annotation.
3. Predicting Disease Variants Using Deep Learning
Predicting disease variants is a critical step in understanding the genetic basis of disease. Deep learning models can prioritize and filter genetic variants to identify those that are most likely to be pathogenic.
3.1. The Process of Variant Prioritization
Variant prioritization is the process of determining the most likely pathogenic variant within genetic screening that damages gene function and underlies the disease phenotype. This process typically involves:
- Variant annotation: Discovering clinically insignificant variants, such as synonymous, deep-intronic variants, and benign polymorphisms.
- Filtering: Removing the clinically insignificant variants to focus on the remaining variants, such as known variants or variants of unknown clinical significance (VUSs).
- Prioritization: Ranking the remaining variants based on their likelihood of being pathogenic.
Variant prioritization is crucial for reducing the number of candidate variants and focusing on those that are most likely to be disease-causing.
3.2. Limitations of Traditional Annotation Approaches
Traditional annotation approaches, such as PolyPhen, SIFT, and GERP, have limitations:
- Rarity: Difficulties in interpreting rare genetic variants in individuals and understanding their impacts on disorder risk.
- Sample size: Statistical methods, such as GWAS, require heavy sampling to distinguish rare genetic variants.
- De novo variants: Traditional approaches cannot deliver information about de novo variants.
These limitations highlight the need for improved variant prioritization methods.
3.3. Deep Learning Models for Prioritizing Variants
Deep learning models have emerged as a powerful method for prioritizing variants. By leveraging the deep neural network (DNN) architecture, these models can learn complex patterns and relationships that are not apparent to traditional methods.
Some notable deep learning models for prioritizing variants include:
- Basset: Basset relies on a CNN algorithm and is designed to predict the causative SNP exploiting DNase I hypersensitivity sequencing data as an input.
- DeepWAS: DeepWAS relies on a CNN algorithm that allows regulatory impact prediction of each variant on numerous cell-type-specific chromatin features.
- DeepPVP: DeepPVP is designed for phenotype-based prioritization of causative variants using deep learning.
3.4. Advantages of Deep Learning in Variant Prediction
Deep learning offers several advantages for predicting disease variants:
- Improved accuracy: Deep learning models can achieve higher accuracy in predicting disease variants compared to traditional methods.
- Integration of multiple data types: Deep learning models can integrate data from multiple sources, such as genomic data, transcriptomic data, and clinical data.
- Identification of novel genes: Deep learning approaches are particularly suited for variant investigation for genes not yet related to specific disease phenotypes.
- Decreased waiting times: Deep learning models can contribute to decreasing waiting times for results and can prioritize variants for further functional analysis.
3.5. Practical Applications
The practical applications of deep learning in disease variant prediction are vast and include:
- Rare disease diagnosis: Identifying disease-causing variants in individuals with rare genetic disorders.
- Cancer genomics: Prioritizing somatic mutations in cancer genomes to identify driver mutations and therapeutic targets.
- Personalized medicine: Predicting an individual’s risk of developing a disease based on their genetic makeup.
LEARNS.EDU.VN offers training and resources to assist in the application of deep learning for disease variant prediction. Consider exploring our courses for hands-on experience and guidance.
4. Gene Expression and Regulation Insights from Deep Learning
Gene expression and regulation are fundamental processes in biology. Deep learning models are providing new insights into these processes, allowing researchers to understand how genes are turned on and off in different cell types and under different conditions.
4.1. The Complexity of Gene Expression
Gene expression involves the initial transcriptional regulators (e.g., pre-mRNA splicing, transcription, and polyadenylation) to functional protein production. Understanding the quantitative regulation of gene expression is challenging due to the complexity of the underlying biological processes.
Traditional approaches for studying gene expression have limitations:
- Limited exploration: Huge biological sequence regions cannot be explored using experimental or computational techniques.
- Promoter focus: The majority of natural mRNA screening approaches focus on studying promoter regions.
- Data integration: Integrating data from multiple sources, such as chromatin accessibility, ChIP-seq, and DNase-seq, is challenging.
4.2. Single-Cell RNA Sequencing (scRNA-seq)
Single-cell RNA sequencing (scRNA-seq) is a powerful technology that allows researchers to measure gene expression levels in individual cells. This technology provides valuable information into cellular heterogeneity that could expand the interpretation of human diseases and biology.
Key applications of scRNA-seq data understanding include:
- Detecting the type and state of the cells.
- Clustering the data.
- Retrieving the data.
4.3. Deep Learning for Predicting Regulatory Sequence Elements
Deep learning has empowered essential progress for constructing predictive methods linking regulatory sequence elements to the molecular phenotypes. By learning complex patterns in the data, deep learning models can predict the effects of regulatory sequence elements on gene expression.
Some notable deep learning models for predicting regulatory sequence elements include:
- SpliceAI: SpliceAI is a deep residual neural network that predicts splice function using only pre-mRNA transcript sequencing as inputs.
- MPRA-DragoNN: MPRA-DragoNN is based on CNN architecture for prediction and analysis of the transcription regulatory activity of non-coding DNA sequencing data measured from (MPRAs) data.
- Xpresso: Xpresso is a deep convolutional neural network (CNN) that conjointly models the promoter sequence and its related mRNA stability features to predict the gene expression levels of mRNA.
- DARTS: DARTS is a deep learning-based model that uses a wide-ranging RNA-seq resources of a various alternative splicing.
- COSSMO: COSSMO adapts to various quantities of alternative splice sites and precisely estimates them via genome-wide cross-validation.
4.4. Applications in Gene Regulation
Deep learning models have a wide range of applications in gene regulation:
- Predicting splicing: Identifying how pre-mRNA transcripts are spliced to produce different mRNA isoforms.
- Analyzing regulatory activity: Predicting the transcription regulatory activity of non-coding DNA sequences.
- Modeling gene expression levels: Predicting gene expression levels based on promoter sequence and mRNA stability features.
- Understanding heterogeneous datasets: Discovering automated relationships among heterogeneous datasets in imperfect biological situations.
4.5. Advancing Gene Expression Studies with Deep Learning
Deep learning is advancing gene expression studies by providing:
- Improved accuracy: Deep learning models can achieve higher accuracy in predicting gene expression levels compared to traditional methods.
- Integration of multiple data types: Deep learning models can integrate data from multiple sources, such as genomic data, transcriptomic data, and epigenetic data.
- Identification of novel regulatory elements: Deep learning models can identify novel regulatory elements that are not apparent to traditional methods.
At LEARNS.EDU.VN, you can find comprehensive resources on deep learning applications in gene expression. Enhance your skills and stay updated with the latest advancements.
 Gene expression regulation
Gene expression regulation
5. Epigenomics and Deep Learning: Unraveling the Code
Epigenomics, the study of heritable changes in gene expression that do not involve alterations to the DNA sequence, is another area where deep learning is making significant contributions. Deep learning models can predict epigenetic marks and their effects on gene expression, offering new insights into the regulation of the genome.
5.1. The Role of Epigenetics
Epigenetic mechanisms, including DNA methylation, histone modifications, and non-coding RNAs, are considered fundamental in understanding disease developments and finding new treatment targets. These mechanisms play a crucial role in regulating gene expression and influencing phenotype.
Challenges in epigenomics include:
- Data interpretation: Complications in developing data interpretation tools to advances in next-generation sequencing and microarray technology to produce epigenetic data.
- Mark interactions: Insufficiency of suitable and efficient computational approaches has led current research to focus on a specific epigenetic mark separately, although several mark interactions and genotypes occurred in vivo.
- Clinical implementation: Epigenetics has yet to be completely employed in clinical implementations.
5.2. Deep Learning Models for Epigenomic Analysis
Deep learning models have achieved success in predicting 3D chromatin interactions, methylation status from single-cell datasets, and histone modification sites based on DNase-Seq data.
Some notable deep learning models for epigenomic analysis include:
- DeepCpG: Accurate prediction of single-cell DNA methylation states using deep learning.
- DeepHistone: A deep learning approach to predicting histone modifications.
- Deopen: A hybrid deep CNN model applied to predict chromatin accessibility within a whole genome from learned regulatory DNA sequence codes.
5.3. Applications in Epigenomics Research
Deep learning models have a wide range of applications in epigenomics research:
- Predicting chromatin accessibility: Predicting chromatin accessibility within a whole genome from learned regulatory DNA sequence codes.
- Predicting histone modifications: Predicting histone modifications to various site-specific markers.
- Discriminating functional SNPs: Revealing the capability to discriminate functional SNPs from their adjacent genetic variants.
5.4. The Power of Deep Learning in Epigenomics
Deep learning provides several advantages for epigenomic analysis:
- Improved accuracy: Deep learning models can achieve higher accuracy in predicting epigenetic marks and their effects on gene expression compared to traditional methods.
- Integration of multiple data types: Deep learning models can integrate data from multiple sources, such as genomic data, transcriptomic data, and epigenetic data.
- Identification of novel epigenetic marks: Deep learning models can identify novel epigenetic marks that are not apparent to traditional methods.
5.5. Resources for Deep Learning in Epigenomics
LEARNS.EDU.VN provides resources and education to help researchers and students leverage the power of deep learning in epigenomics. Our courses offer hands-on experience with the latest tools and techniques.
6. Pharmacogenomics and Deep Learning: Tailoring Treatments
Pharmacogenomics, the study of how genes affect a person’s response to drugs, is another area where deep learning is proving to be valuable. Deep learning models can predict drug response based on an individual’s genetic makeup, helping to tailor treatments and improve patient outcomes.
6.1. Understanding Pharmacogenomics
Pharmacogenomics involves knowledge concerning the association between genetic variants in enormous gene clusters up to whole genomes and the impacts of varying drugs. Understanding the underlying mechanisms of variability in medication response is a key challenge in modern therapeutic methods.
Challenges in pharmacogenomics include:
- Variability: Understanding the underlying mechanisms of variability in medication response.
- Clinical experiments: The clinical experiments generate various errors during the investigation of drug combination efficiency, which is time- and cost-intensive.
- Limitations of traditional approaches: Approaches are limited to particular targets, pathways, or certain cell lines and sometimes need a particular omics dataset of treated cell lines with specific compounds.
6.2. Deep Learning Models for Predicting Drug Response
Deep learning models are reportedly well suited to treatment response prediction tasks based on cell-line omics datasets. By learning complex patterns in the data, these models can predict how an individual will respond to a particular drug.
Some notable deep learning models for predicting drug response include:
- DrugCell: A visible neural network (VNN) interpretation model for the structure and function of human cancer cells in therapy response.
- DeepBL: Based on deep learning architecture executed based on Small VGGNet structure (a type of CNNs) and TensorFlow library to detect beta-lactamases (BLs).
- DeepSynergy: Predicts anti-cancer drug synergy with deep learning.
6.3. Statistical Analysis in Pharmacogenomics
Statistical methods, such as the analysis of variance (ANOVA) test, are utilized to investigate pharmacogenomics associations. This can identify, for example, oncogenic changes that occur in patients, which are indicators of drug-sensitivity variances in cell lines.
To move beyond the drug’s relations to the actual drug reaction predictions, numerous statistical and machine learning methods can be employed, from linear regression models to nonlinear ones, such as kernel methods, neural networks, and SVM.
6.4. Applications in Personalized Medicine
Deep learning models have a wide range of applications in personalized medicine:
- Predicting drug response: Predicting an individual’s response to a particular drug based on their genetic makeup.
- Tailoring treatments: Tailoring treatments to an individual’s specific genetic profile.
- Improving patient outcomes: Improving patient outcomes by selecting the most effective treatments.
6.5. Benefits of Deep Learning in Pharmacogenomics
Deep learning offers several benefits for pharmacogenomics research:
- Improved accuracy: Deep learning models can achieve higher accuracy in predicting drug response compared to traditional methods.
- Integration of multiple data types: Deep learning models can integrate data from multiple sources, such as genomic data, transcriptomic data, and clinical data.
- Identification of novel drug targets: Deep learning models can identify novel drug targets that are not apparent to traditional methods.
- Reduced clinical trial costs: Deep learning models can help to reduce the costs of clinical trials by identifying patients who are most likely to respond to a particular drug.
LEARNS.EDU.VN is dedicated to helping you master the applications of deep learning in pharmacogenomics. Explore our courses to begin your journey.
7. The Future of Deep Learning in Genomics
The field of deep learning in genomics is rapidly evolving, with new algorithms, tools, and applications emerging all the time. As the cost of sequencing continues to decrease and the amount of genomic data continues to grow, deep learning is poised to play an increasingly important role in advancing our understanding of biology and disease.
7.1. Emerging Trends and Technologies
Several emerging trends and technologies are shaping the future of deep learning in genomics:
- Graph neural networks (GNNs): GNNs are designed to analyze data that can be represented as graphs, such as biological networks. These models can capture complex relationships between genes, proteins, and other biological entities.
- Attention mechanisms: Attention mechanisms allow deep learning models to focus on the most important parts of the input data. This can improve the accuracy and interpretability of the models.
- Explainable AI (XAI): XAI techniques are being developed to make deep learning models more transparent and interpretable. This is crucial for building trust in these models and ensuring that they are used responsibly.
- Federated learning: Federated learning allows deep learning models to be trained on decentralized data sources without sharing the data directly. This can enable researchers to collaborate and share insights while protecting patient privacy.
7.2. Overcoming Challenges
Several challenges remain in the field of deep learning in genomics:
- Data quality: Genomic data can be noisy and contain errors, which can negatively impact the performance of deep learning models.
- Interpretability: Deep learning models can be difficult to interpret, making it challenging to understand why they make certain predictions.
- Generalizability: Deep learning models may not generalize well to new datasets or populations.
- Computational resources: Training deep learning models can require significant computational resources, limiting accessibility for some researchers.
Addressing these challenges will be critical for realizing the full potential of deep learning in genomics.
7.3. Ethical Considerations
The use of deep learning in genomics raises several ethical considerations:
- Privacy: Genomic data is highly sensitive and must be protected from unauthorized access.
- Bias: Deep learning models can perpetuate and amplify biases in the data, leading to unfair or discriminatory outcomes.
- Transparency: It is important to be transparent about the limitations of deep learning models and the potential for errors.
- Equity: It is important to ensure that the benefits of deep learning in genomics are shared equitably across all populations.
Addressing these ethical considerations will be crucial for ensuring that deep learning is used responsibly and ethically in genomics.
7.4. Transformative Potential
Despite these challenges, deep learning has the potential to transform genomics in several ways:
- Accelerating discovery: Deep learning can accelerate the discovery of new disease-causing genes and therapeutic targets.
- Improving diagnostics: Deep learning can improve the accuracy and sensitivity of diagnostic tests.
- Personalizing medicine: Deep learning can help to personalize treatments based on an individual’s genetic makeup.
- Advancing our understanding of biology: Deep learning can provide new insights into the complex biological processes that govern life.
7.5. Continuing Education and Resources
LEARNS.EDU.VN is committed to providing continuing education and resources to help researchers and students stay up-to-date on the latest advances in deep learning in genomics. We offer a variety of courses, tutorials, and datasets to help you learn the skills you need to succeed in this exciting field.
With the right knowledge and tools, you can contribute to the transformative potential of deep learning in genomics.
8. Practical Implementation and Tools
To harness the power of deep learning in genomics, researchers and students need access to practical tools and implementation strategies. This section provides an overview of the tools, libraries, and platforms that are commonly used in the field.
8.1. Essential Deep Learning Libraries
Several deep learning libraries are essential for implementing deep learning models in genomics:
- TensorFlow: TensorFlow is a popular open-source deep learning library developed by Google. It provides a flexible and scalable platform for building and training deep learning models.
- Keras: Keras is a high-level API for building and training deep learning models. It is designed to be user-friendly and easy to learn, making it a good choice for beginners.
- PyTorch: PyTorch is another popular open-source deep learning library developed by Facebook. It is known for its flexibility and dynamic computational graph, making it a good choice for research.
- scikit-learn: scikit-learn is a popular machine learning library that provides a wide range of algorithms for classification, regression, and clustering. It is often used for preprocessing data and evaluating the performance of deep learning models.
8.2. Genomic Data Formats and Handling
Handling genomic data requires familiarity with specific data formats and tools:
- FASTQ: FASTQ is a common format for storing DNA and RNA sequences. It contains the sequence reads and their associated quality scores.
- SAM/BAM: SAM (Sequence Alignment/Map) and BAM (Binary Alignment/Map) are formats for storing aligned sequencing reads. They contain information about the alignment of each read to a reference genome.
- VCF: VCF (Variant Call Format) is a format for storing genetic variants. It contains information about the location, type, and frequency of each variant.
- BED: BED (Browser Extensible Data) is a format for storing genomic annotations. It contains information about the location and attributes of genomic features, such as genes, exons, and regulatory elements.
8.3. Cloud Computing Platforms
Cloud computing platforms provide access to the computational resources needed to train deep learning models on large genomic datasets:
- Amazon Web Services (AWS): AWS provides a wide range of cloud computing services, including virtual machines, storage, and databases. It also offers specialized services for genomics, such as Amazon Genomics CLI.
- Google Cloud Platform (GCP): GCP provides a similar set of cloud computing services to AWS. It also offers specialized services for genomics, such as Google Cloud Life Sciences.
- Microsoft Azure: Microsoft Azure is another popular cloud computing platform. It also offers specialized services for genomics, such as Azure Genomics.
8.4. Example Workflow for Variant Calling
Here is an example workflow for variant calling using deep learning:
- Data acquisition: Obtain FASTQ files from a sequencing facility.
- Data preprocessing: Preprocess the FASTQ files to remove errors and biases.
- Alignment: Align the sequencing reads to a reference genome using a tool like BWA or Bowtie2.
- Variant calling: Call variants using a deep learning model like DeepVariant or Clairvoyante.
- Annotation: Annotate the variants using a tool like ANNOVAR or VEP.
- Filtering: Filter the variants to remove false positives.
- Visualization: Visualize the variants using a tool like IGV or UCSC Genome Browser.
8.5. Best Practices for Implementation
Here are some best practices for implementing deep learning in genomics:
- Data quality: Ensure that the data is of high quality and has been properly preprocessed.
- Model selection: Choose an appropriate deep learning model for the task.
- Hyperparameter tuning: Tune the hyperparameters of the deep learning model to optimize performance.
- Validation: Validate the performance of the deep learning model on an independent dataset.
- Interpretability: Use XAI techniques to make the deep learning model more interpretable.
- Reproducibility: Ensure that the results are reproducible by documenting the workflow and using version control.
LEARNS.EDU.VN provides comprehensive resources and courses that cover these practical implementation aspects, ensuring you are well-equipped to apply deep learning in your genomic research.
9. Case Studies: Deep Learning Success Stories in Genomics
To illustrate the transformative potential of deep learning in genomics, let’s examine some real-world case studies where deep learning has led to significant breakthroughs.
9.1. Case Study 1: Deep Learning for Cancer Diagnosis
- Problem: Traditional methods for diagnosing cancer can be slow and inaccurate, leading to delays in treatment.
- Solution: Researchers developed a deep learning model that can diagnose cancer from medical images with high accuracy.
- Results: The deep learning model achieved higher accuracy than human experts and reduced the time required for diagnosis.
- Impact: Improved cancer diagnosis and treatment, leading to better patient outcomes.
9.2. Case Study 2: Deep Learning for Drug Discovery
- Problem: Traditional methods for drug discovery are time-consuming and expensive, with a high failure rate.
- Solution: Researchers developed a deep learning model that can predict the efficacy of drug candidates with high accuracy.
- Results: The deep learning model identified several promising drug candidates that were subsequently validated in preclinical studies.
- Impact: Accelerated drug discovery, leading to the development of new treatments for disease.
9.3. Case Study 3: Deep Learning for Personalized Medicine
- Problem: Traditional treatments are often ineffective because they do not take into account individual genetic differences.
- Solution: Researchers developed a deep learning model that can predict an individual’s response to a particular treatment based on their genetic makeup.
- Results: The deep learning model identified the most effective treatments for each individual, leading to improved patient outcomes.
- Impact: Personalized medicine, leading to more effective and targeted treatments.
9.4. Specific Example: Predicting Cancer Drug Response
A study used deep learning to predict how cancer cells would respond to various drugs based on genomic and transcriptomic data. The model integrated data from multiple sources to predict drug sensitivity, outperforming traditional methods and identifying potential biomarkers for drug response.
9.5. Success in Identifying Genetic Variants
Deep learning models have been successfully used to identify genetic variants associated with Alzheimer’s disease. By analyzing large-scale genomic data, these models have pinpointed novel genetic markers that contribute to disease risk.
9.6. Success in Enhancing Diagnostics
Deep learning algorithms are being employed to enhance the accuracy and speed of diagnostic tests for various diseases. For example, they can analyze medical images to detect early signs of diseases with greater precision than traditional methods.
LEARNS.EDU.VN provides resources and courses that highlight these case studies, demonstrating the real-world impact and application of deep learning in genomics.
10. Frequently Asked Questions (FAQ)
Here are some frequently asked questions about deep learning in genomics:
- What is deep learning in genomics?
- Deep learning in genomics is the application of deep learning algorithms to analyze and interpret genomic data.
- What types of genomic data can be analyzed using deep learning?
- Deep learning can be used to analyze a wide range of genomic data, including DNA sequences, RNA sequences, gene expression levels, and epigenetic marks.
- What are the benefits of using deep learning in genomics?
- Deep learning can improve the accuracy and sensitivity of genomic analysis, accelerate discovery, and personalize medicine.
- What are some common deep learning algorithms used in genomics?
- Common deep learning algorithms used in genomics include CNNs, RNNs, and DNNs.
- What are some challenges in using deep learning in genomics?
- Challenges include data quality, interpretability, generalizability, and computational resources.
- What ethical considerations should be taken into account when using deep learning in genomics?
- Ethical considerations include privacy, bias, transparency, and equity.
- How can I get started with deep learning in genomics?
- You can get started by taking courses, reading tutorials, and experimenting with open-source tools and datasets.
- What are some cloud computing platforms that can be used for deep learning in genomics?
- Cloud computing platforms include AWS, GCP, and Microsoft Azure.
- What are some essential deep learning libraries for genomics?
- Essential deep learning libraries include TensorFlow, Keras, PyTorch, and scikit-learn.
- Where can I find more information about deep learning in genomics?
- You can find more information at LEARNS.EDU.VN and other online resources.
Conclusion: Embracing Deep Learning in Genomics
Deep learning is revolutionizing genomics, offering unprecedented opportunities for understanding and manipulating the building blocks of life. From enhancing variant calling to predicting drug responses, the applications of deep learning in genomics are vast and transformative. At LEARNS.EDU.VN, we are committed to providing the resources and education you need to harness the power of deep learning and contribute to this exciting field.
We encourage you to explore the resources and courses available at LEARNS.EDU.VN to further your understanding and skills in deep learning and genomics. Our comprehensive materials, hands-on projects, and expert guidance will help you stay ahead in this rapidly evolving field.
Ready to dive deeper? Visit LEARNS.EDU.VN today to explore our courses and resources!
Contact Information:
- Address: 123 Education Way, Learnville, CA 90210, United States
- WhatsApp: +1 555-555-1212
- Website: LEARNS.EDU.VN
Take the next step in your education journey with learns.edu.vn and unlock the full potential of deep learning in genomics.